Otsi
Otsi
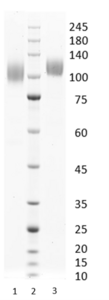
Figure 1. Simply Blue Safe stained SDS-PAGE analysis of SARS-CoV-2 Spike S1 Stealth Omicron (BA.2). 4-12% gradient gel is used for analysis. Lane 1. 0.8 µg SARS-CoV-2 Spike S1 Stealth Omicron (BA.2) (-DTT). Lane 2. Protein marker (Smobio). Lane 3. 0.8 µg SARS-CoV-2 Spike S1 Stealth Omicron (BA.2) (+DTT).
Figure 2. HPLC analytical SEC for final product.
Figure 3. HPLC analytical SEC after 3 freeze-thaw cycles.
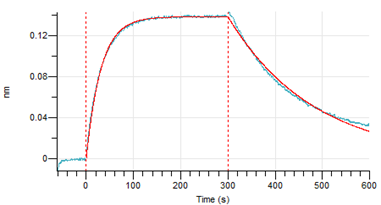 Figure 4. Octet RED96e analysis of SARS-CoV-2 S1 Omicron protein binding to the ACE2 receptor.
Figure 4. Octet RED96e analysis of SARS-CoV-2 S1 Omicron protein binding to the ACE2 receptor.
Catalogue #
P-375-100
Name
SARS-CoV-2 Spike S1 Stealth Omicron (BA.2)
Description
Protein contains amino acids 14-681, mutations T19I, L24S, del25/27, G142D, V213G, G339D, S371F, S373P, S375F, T376A, D405N, R408S, K417N, N440K, S477N, T478K, E484A, Q493R, Q498R, N501Y, Y505H, D614G, H655Y, N679K, P681H, two extra amino acids (AS) in N-terminus, His-6 tag at C-terminus and GSG linker between protein and tag.
Uniprot ID
P0DTC2
MW
75.81 kDa
Host
CHO-based cell line (expressed by QMCF Technology)
Purification
Purified by Ni-affinity chromatography and gel-filtration from serum-free CHO growth media, sterile filtrated
Concentration
1 mg/ml
Buffer
PBS pH 7.4
Endotoxine
NA
Bioproperties
Measured by its binding ability to ACE2 protein by OCTET RED96 system.
QC
SDS-PAGE, NanoDrop, A280 Analytical SEC, Octet binding to ACE2
Shipping
Shipped on dry ice.
Storage
Store at -70°C upon receipt. Recommended to aliquot into smaller quantities. Avoid repeated freeze-thaw cycles
This product is for research use only
TECHNICAL ASSISTANCE
Please refer any technical questions to
technical.support@icosagen.com